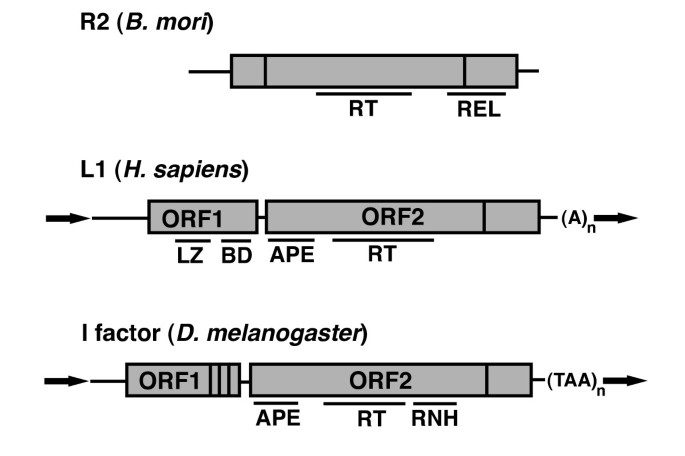
Non-long terminal repeat (non-LTR) retrotransposons: mechanisms, recent developments, and unanswered questions | Mobile DNA | Full Text

Estimation of long terminal repeat element content in the Helicoverpa zea genome from high-throughput sequencing of bacterial artificial chromosome pools

Sensitive detection of pre-integration intermediates of long terminal repeat retrotransposons in crop plants | Nature Plants

Identification of DNA Sequences of Long Terminal Repeat Retrotransposons in Maize — Journal of Young Investigators

Structure of newly identified long terminal repeat retrotransposons... | Download Scientific Diagram

An endogenous retroviral long terminal repeat is the dominant promoter for human β1,3-galactosyltransferase 5 in the colon | PNAS
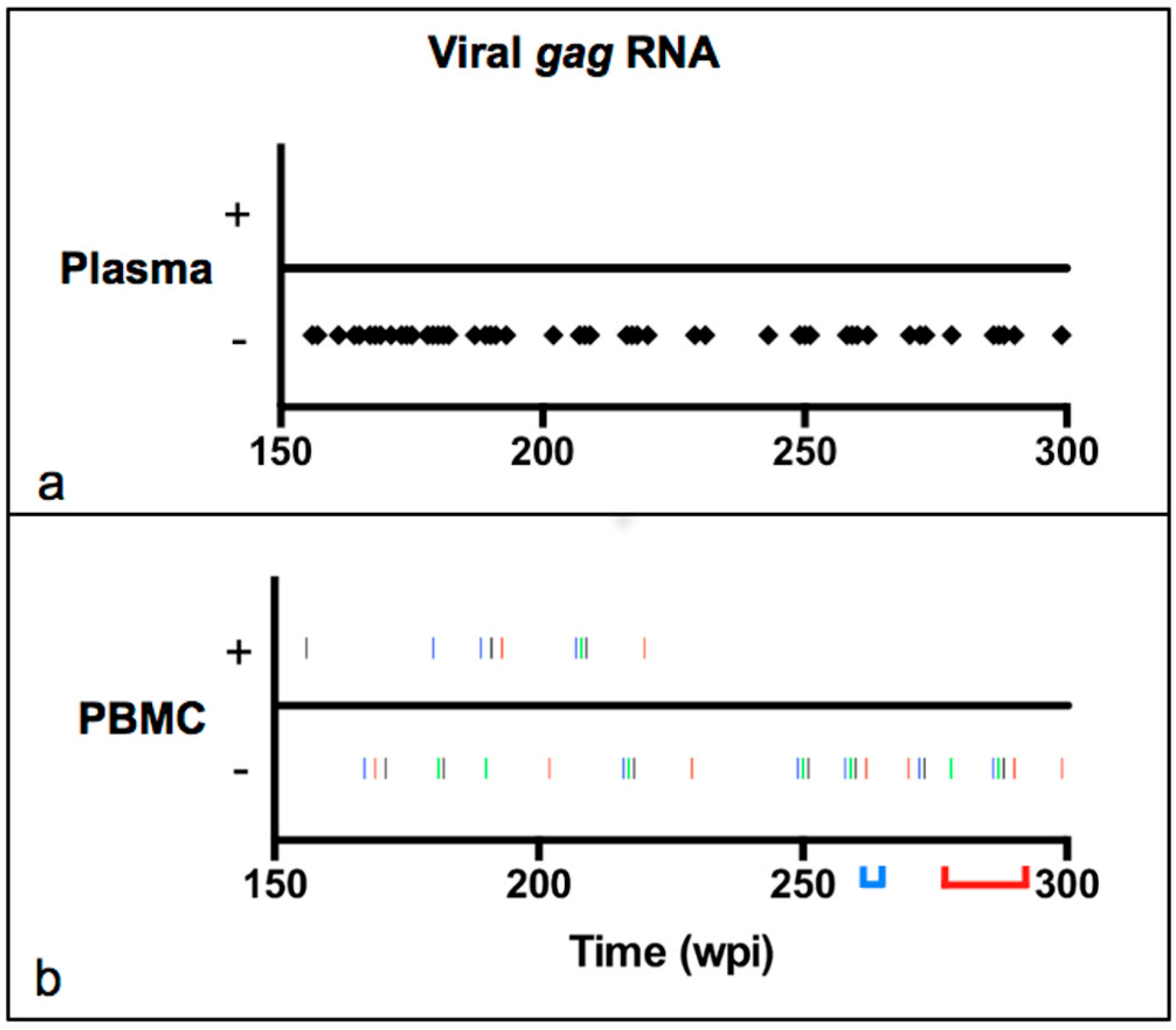
Veterinary Sciences | Free Full-Text | Sequence Instability in the Proviral Long Terminal Repeat and gag Regions from Peripheral Blood and Tissue-Derived Leukocytes of FIV-Infected Cats during the Late Asymptomatic Phase
NFAT5 Regulates HIV-1 in Primary Monocytes via a Highly Conserved Long Terminal Repeat Site | PLOS Pathogens

Viruses | Free Full-Text | Genetic Diversity in HIV-1 Subtype C LTR from Brazil and Mozambique Generates New Transcription Factor-Binding Sites

Global long terminal repeat activation participates in establishing the unique gene expression programme of classical Hodgkin lymphoma | Leukemia